기술동향
Shaping the matrix
- 등록일2009-06-08
- 조회수7265
- 분류기술동향
-
자료발간일
2009-06-05
-
출처
RIKEN Research
- 원문링크
-
키워드
#시냅스#synaptic# synaptic scaffolding proteins
출처 : RIKEN Research
Shaping the matrix
A mesh-like structure formed by two synaptic scaffolding proteins controls the shape and protein make-up of the synapse
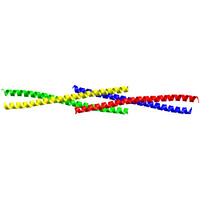
Figure 1: When four Homer protein monomers interact, they form a long tail-to-tail tetramer. The crystal structure of the carboxy-terminal of Homer is shown; each of the four Homer monomers is represented by a different color.
An international team of researchers, including Mariko Kato Hayashi at the RIKEN Brain Science Institute in Wako, has found that the scaffolding proteins Shank and Homer polymerize to form a large matrix that regulates key features of the synapse1-namely its shape, protein composition, and function.
Chemical communication between neurons occurs at the synapse. Many synaptic proteins-such as neurotransmitter receptors, scaffolding proteins, and signaling proteins-are poised at the receiving end of the synapse, called the ‘postsynaptic density’ (PSD). Increasing the expression of the PSD scaffolding proteins Shank and Homer within neurons increases the size of dendritic spines, in which the PSD is located. These new findings explain why.
The researchers incubated pure Shank protein, pure Homer protein, and a mixture of both proteins to allow them to interact. When they observed these three different preparations under the electron microscope, the pure Shank protein looked like compact globs, while the pure Homer protein looked like longer threads.
Notably, incubation of both Shank and Homer together resulted in a large mesh-like structure, with Shank proteins forming ‘hubs’ which attached to the ends of Homer protein fibers. The formation of this large Homer?Shank matrix within neurons may explain why the dendritic spines of these neurons are abnormally large when these two proteins are artificially expressed at high levels.
Using a technique called x-ray crystallography, which allows scientists to determine the shape of proteins, the researchers were able to establish the structural basis by which Homer forms long filaments: four Homer protein monomers interact in a special crossing tail-to-tail tetrad structure (Fig. 1). Interference with the creation of this structure, by mutating key parts of the tetrad structure, blocks formation of the Homer?Shank matrix.
The team found that expressing this mutant Homer in neurons resulted in abnormal synaptic development and the normal accumulation of Shank and other synaptic proteins was reduced at the synapse. Also, the dendritic spines of these neurons were long and thin, which is characteristic of immature spines. These s seemed to affect neuronal function, because neurons expressing the mutant Homer protein, which could not form the tetrad structure, had reduced excitatory neurotransmission compared with neurons expressing the normal Homer protein.
“The Homer?Shank matrix could be the structural framework of the PSD,” says Hayashi, “and could work as the platform for other important synaptic proteins. It would therefore be playing a role much like the circuit board of a computer.”
Reference
1. Hayashi, M.K., Tang, C., Verpelli, C., Narayanan, R., Stearns, M.H., Xu, R.-M., Li, H., Sala, C. & Hayashi, Y., The postsynaptic density proteins Homer and Shank form a polymeric network structure. Cell 137, 159?171 (2009). | article |
The corresponding author for this highlight is based at the RIKEN Laboratory for Synaptic Function
...................(계속)
☞ 자세한 내용은 내용바로가기 또는 첨부파일을 이용하시기 바랍니다.
-
다음글
- 다음글이 없습니다.
-
이전글
- 이전글이 없습니다.
관련정보
동향
- 기술동향 [전자통신동향분석] 뉴로모픽 감각 인지 기술 동향―촉각, 후각을 중심으로 2023-12-07
- 기술동향 시냅스 신경과학 최근 연구동향 2022-06-28
- 기술동향 커넥톰 발생의 기전 2021-02-02
- 기술동향 감정 기억의 저장과 회상을 관장하는 편도체(Amygdala)의 신경회로에서 시냅스 가소성(Synaptic Plasticity), 기억 심상(Engram)과 신경망 진동(Network oscillation)의 중요성 2018-11-12
- 기술동향 단백질 3차 구조 기반 시냅스 접착단백질 작용 기전 이해 2018-01-24

